
The cdom package implements various functions used to model and calculate metrics from absorption spectra of chromophotic dissolved organic matter (CDOM).
This package provides:
The package can be installed using the following command.
devtools::install_github("PMassicotte/cdom")Please note that this is a developing version of the package for testing only. Please fill an issue when you find bugs.
All functions from the package start with the cdom_
prefix.
library(cdom)
ls("package:cdom")
## [1] "cdom_exponential" "cdom_slope_ratio" "cdom_spectral_curve"
## [4] "spectra"The cdom_fit_exponential() function fits an exponential
curve to CDOM data using the simple model proposed by Jerlov (1968),
Lundgren (1976), Bricaud, Morel, and Prieur (1981).
a(\lambda) = a(\lambda0)e^{-S(\lambda - \lambda0)} + Klibrary(ggplot2)
library(cdom)
data(spectra)
fit <- cdom_exponential(
wl = spectra$wavelength,
absorbance = spectra$spc3,
wl0 = 350,
startwl = 190,
endwl = 900
)
coef(fit)
## S K a0
## 0.02220677 1.85125088 6.02460462
p <- plot(fit)
## Warning: `aes_string()` was deprecated in ggplot2 3.0.0.
## ℹ Please use tidy evaluation idioms with `aes()`.
## ℹ See also `vignette("ggplot2-in-packages")` for more information.
## ℹ The deprecated feature was likely used in the cdom package.
## Please report the issue at <https://github.com/PMassicotte/cdom/issues>.
## This warning is displayed once per session.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.
p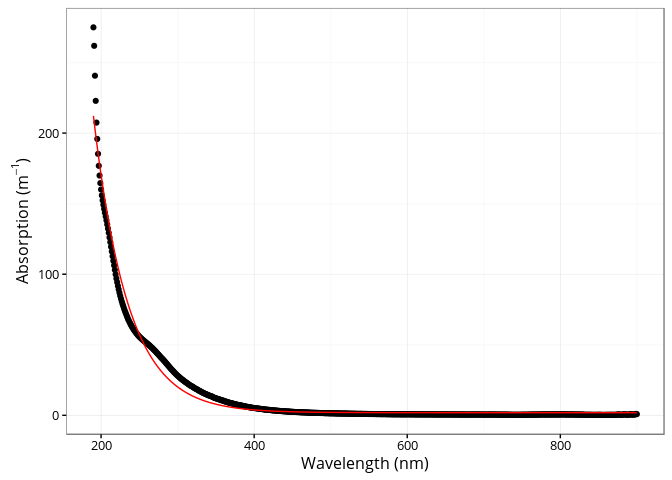
The cdom_slope_ratio() function calculates the slope
ratio (SR) which is defined as:
S275-295/S350-400. See Helms et al. (2008) for
detailed information.
library(cdom)
data(spectra)
cdom_slope_ratio(spectra$wavelength, spectra$spc1)
## [1] 1.325082The cdom_spectral_curve() function generates the
spectral curve using the slope of the linear regression between the
natural log absorption spectrum and wavelengths over a sliding window of
21 nm interval (default) at 1 nm resolution. See Loiselle et al. (2009)
for detailed information.
library(cdom)
data(spectra)
res <- cdom_spectral_curve(
wl = spectra$wavelength,
absorbance = spectra$spc10,
interval = 21,
r2threshold = 0.98
) # Maybe to restrictive...
ggplot(res, aes(x = wl, y = s)) +
geom_point() +
geom_line() +
xlab("Wavelength (nm)") +
ylab(expression(paste(
"Spectral slope (",
nm^{
-1
},
")"
)))
library(dplyr)
##
## Attaching package: 'dplyr'
## The following objects are masked from 'package:stats':
##
## filter, lag
## The following objects are masked from 'package:base':
##
## intersect, setdiff, setequal, union
library(tidyr)
data(spectra)
spectra_nested <- spectra %>%
pivot_longer(
starts_with("spc"),
names_to = "sample",
values_to = "absorption"
) %>%
group_by(sample) %>%
nest() %>%
mutate(
model = purrr::map(
data,
~ cdom_exponential(
.$wavelength,
.$absorption,
wl0 = 350,
startwl = 190,
endwl = 900
)
)
)A total 25 absorption spectra are provided in the package.
library(ggplot2)
library(tidyr)
data(spectra)
spectra <- pivot_longer(
spectra,
starts_with("spc"),
names_to = "sample",
values_to = "absorption"
)
ggplot(spectra, aes(x = wavelength, y = absorption, group = sample)) +
geom_line(size = 0.1) +
xlab("Wavelength (nm)") +
ylab(expression(paste(
"Absorption (",
m^{
-1
},
")"
)))
## Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
## ℹ Please use `linewidth` instead.
## This warning is displayed once per session.
## Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
## generated.
citation("cdom")
## To cite cdom in publications use:
##
## Massicotte, P., and Markager, S. (2016). Using a Gaussian
## decomposition approach to model absorption spectra of chromophoric
## dissolved organic matter. Mar. Chem. 180, 24-32.
## doi:10.1016/j.marchem.2016.01.008.
##
## A BibTeX entry for LaTeX users is
##
## @Article{,
## title = {Using a Gaussian decomposition approach to model absorption spectra of chromophoric dissolved organic matter},
## author = {Philippe Massicotte and Stiig Markager},
## journal = {Marine Chemistry},
## year = {2016},
## volume = {180},
## pages = {24--32},
## url = {https://linkinghub.elsevier.com/retrieve/pii/S0304420316300081},
## }